Projects
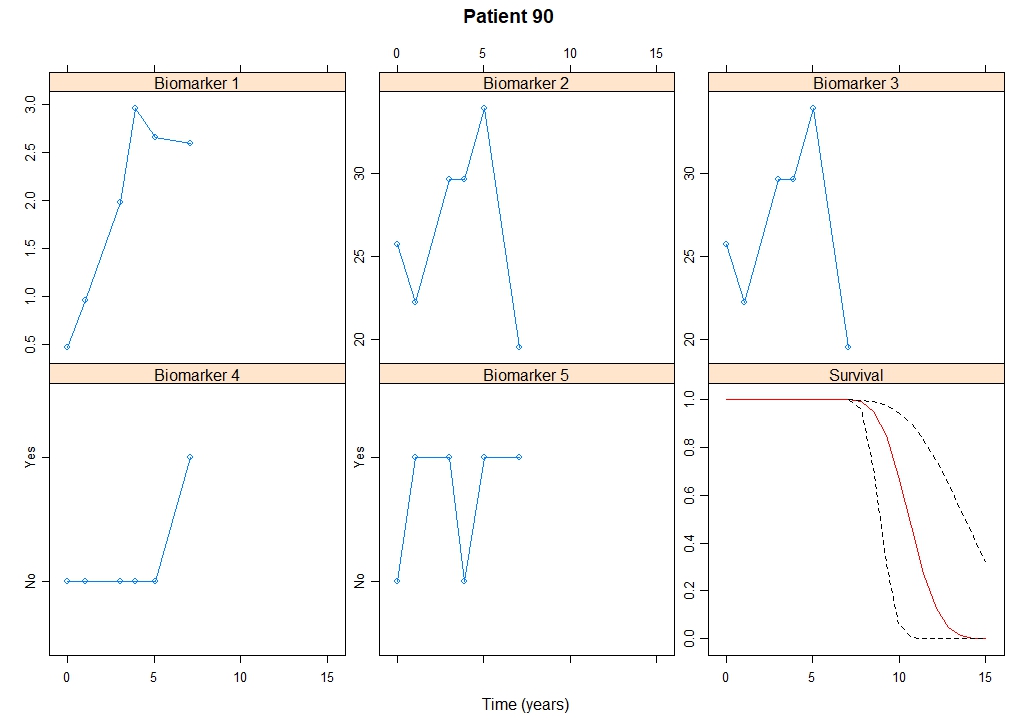
Multivariate Joint Models
Computational methods for multivariate joint models.
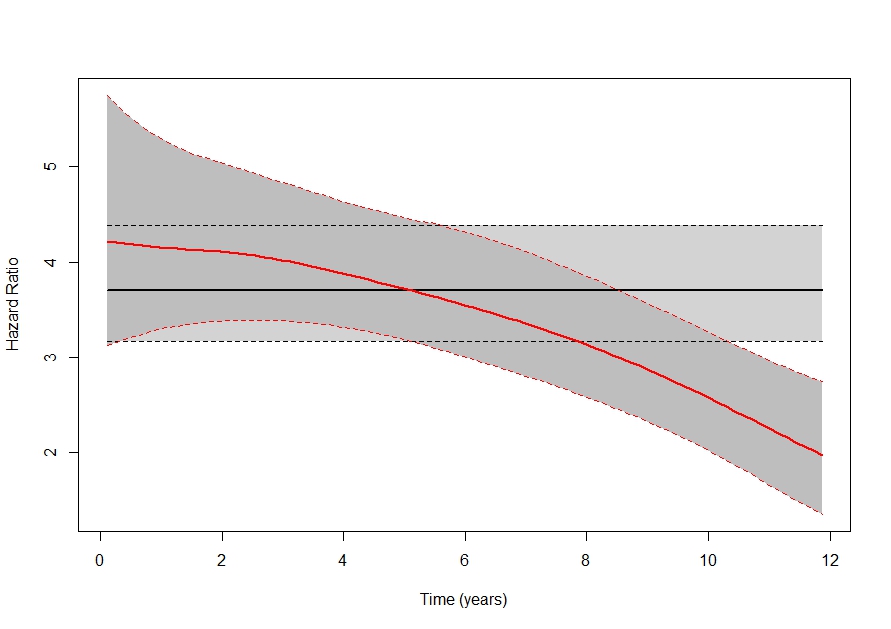
Individualized Predictions, Time-varying Effects and Time-varying Covariates
Extensions of joint models for improving subject-specific predictions.
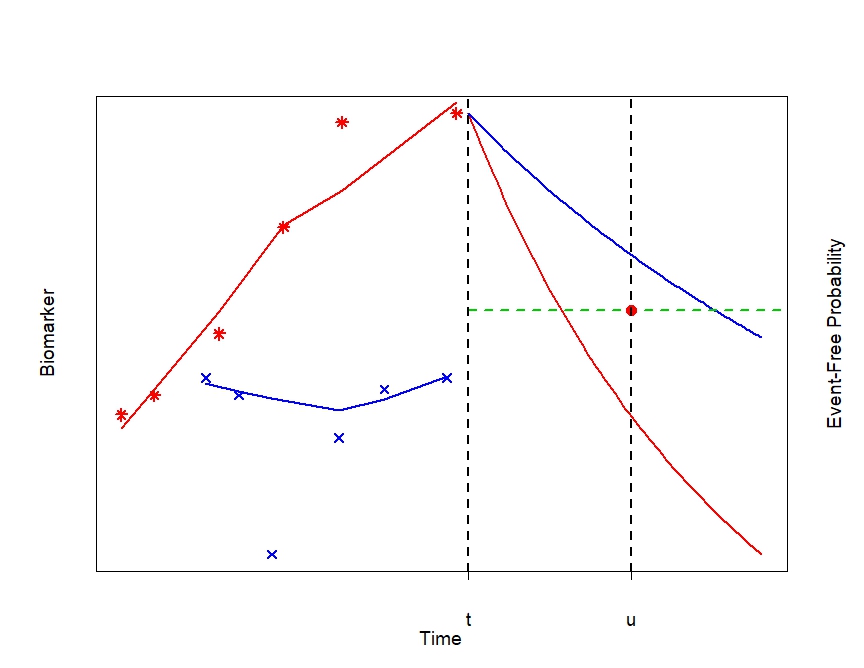
Personalized Active Surveillance and Screening
Novel methods for optimally planning when to collect longitudinal measurements or event information.
